Expanding the genomic landscape of Pseudomonas aeruginosa clone C
Published: 2026-04-23
Pseudomonas aeruginosa is a gram-negative opportunistic pathogen that causes a wide range of infections, including urinary tract infections, and chronic lung infections in cystic fibrosis patients. Beyond clinical settings, it is highly adaptable and persists in diverse environments such as rivers, drinking water systems, and swimming pools. One of its most widespread and successful lineages, clone C (sequence type ST17), is globally distributed across both environmental and clinical niches, and is associated with acute and chronic infections.
To better understand the ecological and pathogenic diversity of clone C, Ham et al.(2026) generated complete genome sequences for three strains isolated from underrepresented environments. Specifically, they sequenced strain 8277 from a urinary tract infection in the United Kingdom, and strains PT31M and SG50M from anthropogenic aquatic environments (drinking and swimming pool water, respectively) in Germany. Prior to this work, complete genomes of clone C strains were largely limited to cystic fibrosis lung isolates, restricting comparative analyses across different ecological niches.
Ham et al.(2026) extracted genomic DNA from cultured cells, and sequenced it using a combination of PacBio SMRT long-read sequencing and Illumina NovaSeq short-read sequencing. Hybrid assembly approaches were applied to generate high-quality genomes, with long reads enabling de novo assembly and short reads used for error correction. The resulting assemblies were polished and annotated using established bioinformatics pipelines, ensuring accurate genome reconstruction and gene prediction.
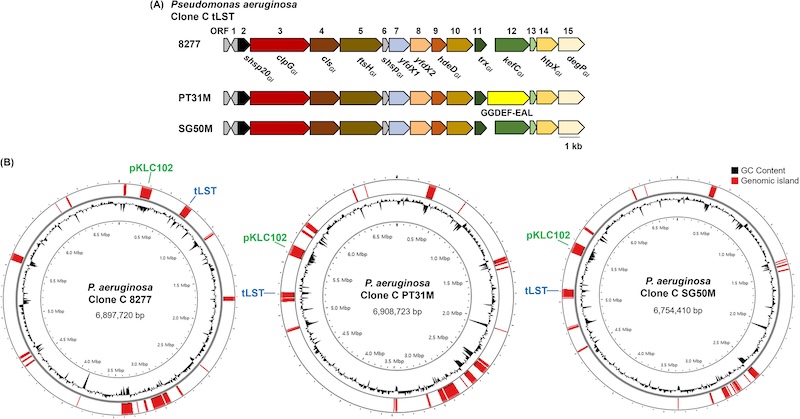
The genomes from all three strains were assembled into single circular chromosomes, with genome sizes ranging from 6.75 to 6.90 Mb and an average GC content of ~66.7%. The genomes contained between 6,156 and 6,318 predicted coding sequences, along with complete sets of rRNA and tRNA genes. Notably, no plasmids were detected. However, the commonly observed clone C mobile element pKLC102 was found integrated into the chromosome in all strains, indicating chromosomal incorporation of mobile genetic elements.
Comparative analysis revealed both shared and distinct genomic features among the strains. All three genomes contained the transmissible locus for stress tolerance (tLST), which encodes proteins such as ClpG disaggregase and small heat shock proteins that enhance bacterial survival under stress conditions. Whilst the tLST region of the genomic islands was highly conserved, others showed variation, reflecting genomic plasticity and adaptation to different environments. For example, strain 8277 showed adaptation to a human urinary tract environment, whereas PT31M and SG50M reflect survival strategies in anthropogenic water systems.
By expanding the number of fully sequenced clone C genomes from diverse sources, Ham et al.(2026) provide valuable reference data for studying the ecological versatility and global success of P. aeruginosa. Their findings highlight how genomic plasticity, combined with conserved stress tolerance mechanisms, enables clone C strains to thrive across both environmental reservoirs and human hosts. This work underscores the importance of including non-clinical isolates in genomic studies to better understand the transmission, persistence, and pathogenic potential of opportunistic bacteria.
Data
- Whole-genome data have been deposited at GenBank under the accession numbers CM129832 (PT31M), CM129833 (SG50M), and CP170200 (8277).
- The associated BioProject and BioSample accession numbers are PRJNA1321591 and SAMN51219634 (PT31M), PRJNA1321598 and SAMN51219831 (SG50M), and PRJNA1162667 and SAMN43818467 (8277), respectively.
- The PacBio and Illumina sequencing raw data for the whole-genome data have been deposited in the NCBI Sequence Read Archive database under the accession numbers SRX30956583 and SRX31989920 (PT31M), SRX30956584 and SRX32015471 (SG50M), and SRX30950279 and SRX32015960 (8277), respectively.
Article
DOI: 10.1128/mra.01240-25
Ham, S., Römling, U., & Lee, C. (2026). Complete genome sequences of Pseudomonas aeruginosa clone C strains 8277, PT31M, and SG50M isolated from the urinary tract and anthropogenic water environments. Microbiology Resource Announcements, e01240-25.
Funding
Ute Römling received funding from the Swedish Research Council for Medicine and Health (K2012-56X-22034-01-3).