A wider variety of data sources contribute to better spatio-temporal predictions of COVID-19 test positivity
Published: 2022-11-22
Over the last three years, the COVID-19 pandemic has challenged healthcare and societal functions worldwide. The rapid development of vaccines and treatments did help to mitigate the effects, though. The COVID-19 pandemic is currently in a less acute phase in most parts of the world, but pandemic preparedness measures are still sorely needed.
COVID-19 has often been found to cause outbreaks in geographical clusters. This is likely due to transmission in enclosed public spaces or places where social interactions occur. As part of pandemic preparedness efforts, local health authorities need emergency strategies in place to curb future outbreaks. These strategies may include measures such as COVID-19 testing, contact-tracing, and/or communication campaigns. The identification of geographical areas where COVID-19 is spreading is key to implementing these strategies effectively.
To date, spatio-temporal COVID-19 prediction models have primarily focused on anticipating the likely number of subsequent COVID-19 cases. The most common approach has been to use single modelling, where recent data is incorporated into models and predictions are made either by extrapolation using past trajectories or using time-series analysis. Another approach is using ensemble forecasting. The latter has been utilised, with varying success, to model several other infectious diseases, such as dengue fever. It combines two or more distinct prediction model outputs into one using weighted averages. In conclusion, available models have shown varying accuracy and power and they lack good geographical resolution. According to the World Health Organization (WHO), the use of fine geographical resolution is one of the main criteria for ensuring that local testing strategies are effective. Epidemic geospatial predictions to locate local outbreaks are therefore warranted.
In a recent publication in Scientific Reports, researchers from Uppsala University and Science for Life Laboratory (First authors: Vera van Zoest, Georgios Varotsis, and Uwe Menzel; Corresponding author: Vera van Zoest; Principal Investigator: Tove Fall) predicted trends in COVID-19 test positivity, which is an important marker for planning for future local testing capacity and accessibility.
van Zoest, Varotsis, Menzel, and colleagues aimed to develop and evaluate the performance of four different statistical single models and one ensemble model in predicting trends in COVID-19 test positivity. The researchers used information about postal services to divide Uppsala County into 50 service point areas. The reason behind this was that, in Sweden, such service points are often located close to major grocery and retail stores, which are places of contact within a neighborhood. The smallest population in a service point area was approximately one thousand inhabitants, and the largest population included approximately 20,000 inhabitants. Data across one year (June 29, 2020-July 4, 2021) was included in the models, and the researchers used a 1-week prediction window. The collated information included both direct and indirect indicators of transmission (e.g. mobility data, number of calls to the national healthcare advice line, and vaccination coverage) as potential predictors.
All data used in the prediction models was collected as part of the CRUSH Covid initiative, an ongoing collaboration between Uppsala County Council and a multidisciplinary team of researchers from Uppsala University. The project aims to assist local health authorities to mitigate local COVID-19 outbreaks by monitoring how the disease spreads over key geographical areas. The data collected has been used to build a comprehensive database that includes descriptive statistics on each data source. This database is freely available in the CRUSH Covid app and is used by both healthcare authorities and the public. Indeed, data from CRUSH Covid has been an important tool for the local health authorities during the pandemic.
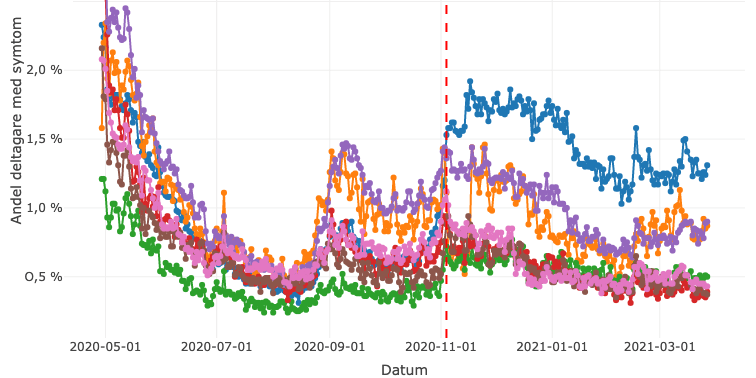
When building the prediction models, the researchers considered a number of factors: (1) the geo-location of the service point areas, (2) demography, (3) direct indicators such as COVID-19 RT-PCR tests performed, (4) weekly hospitalization and ICU beds used, (5) weekly indirect indicators e.g., Health (1177) and 112 emergency numbers assessed as suspected COVID-19, (6) vaccination coverage, and (7) Google mobility data indicating increased social contact. The outcome was set as the prediction of test positivity at each service point, hence the number of COVID-19 tests that were positive as a proportion of all the the tests performed during the following week.
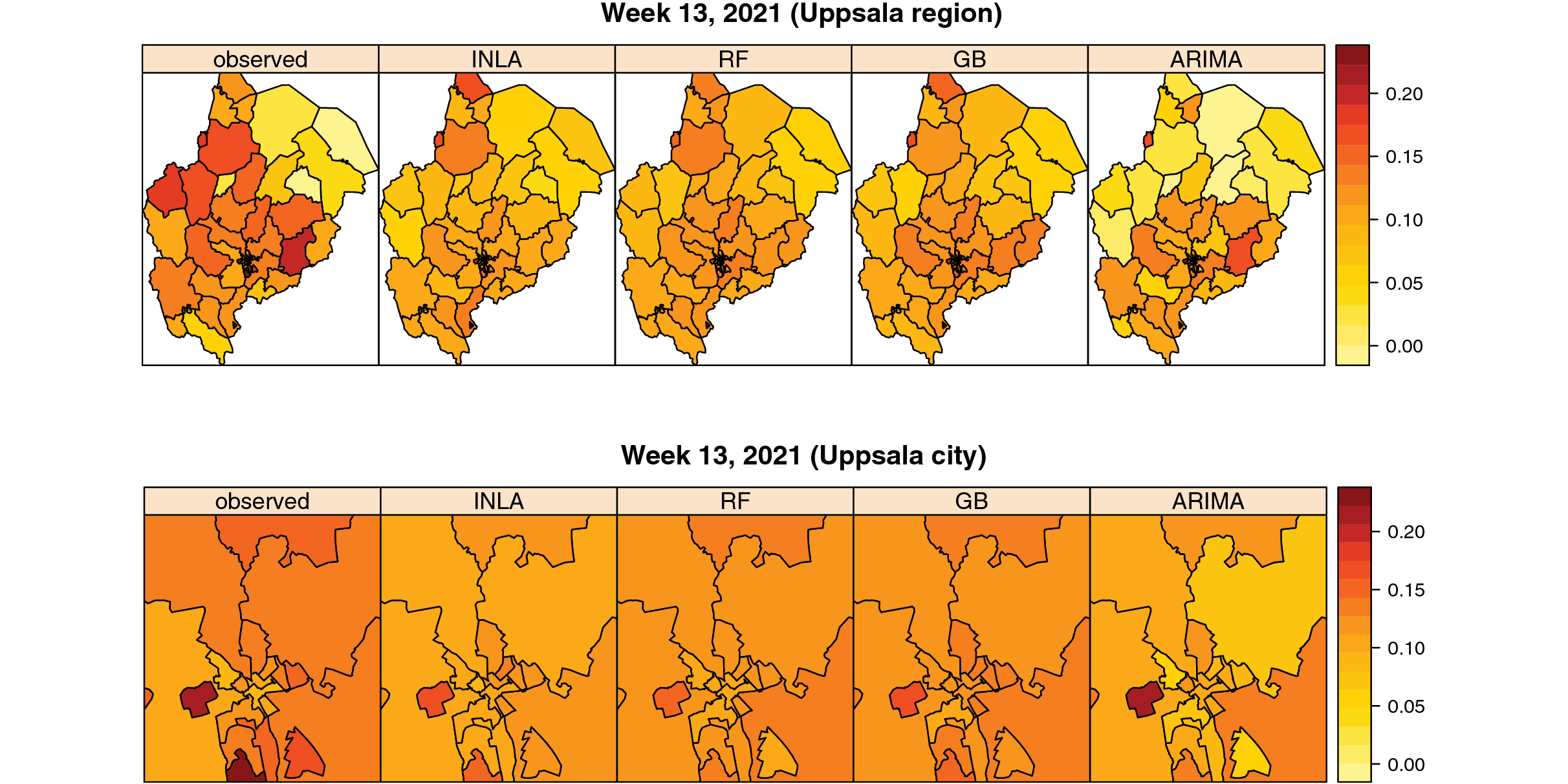
van Zoest, Varotsis and Menzel et al. developed four models for a 1-week-window, using four different statistical methods based on gradient boosting (GB), random forest (RF), autoregressive integrated moving average (ARIMA), and integrated nested Laplace approximations (INLA). These models were built on different selections of covariates. GB and RF models, for example, relied heavily on COVID-19 cases/100,000 inhabitants and health call data. Three of the models (GB, RF and INLA) were found to be better than the naïve baseline model (based on a one-week lagged test positivity), and showed moderate accuracy. The researchers also used an ensemble model, a weighed combination of the three models, which was found to slightly improve accuracy. The researchers found that the accuracy of the models increased over time. This is probably because new data was incorporated weekly from multiple data sources, so the models were gradually based on larger sets of data.
In summary, van Zoest, Varotsis, and Menzel and colleagues’ findings indicate that the collection of data from a wide variety of sources can contribute to better spatio-temporal predictions of COVID-19 test positivity. Combining various data sources into prediction models may prove important in local efforts to curb the spread of COVID-19 and other viral diseases. In the future, complex ensemble models combined with machine learning may provide better and more accurate models to predict the spread of infectious diseases.
Data
In accordance with the principles of open science, the data used in this study is available online at: https://github.com/MolEpicUU/spatiotemporal_predictions_COVID19.
The CRUSH Covid app can be accessed directly for exploration.
The CRUSH Covid project is one of the partner projects featured on the Swedish Pathogens Portal. To read more about this project, please visit the dedicated partner project dashboard.
Funding
Open access funding provided by Uppsala University. The study was partly funded by VINNOVA, the Swedish Research Council, the Swedish Heart-Lung Foundation, and the European Research Council.
Article
DOI: 10.1038/s41598-022-19155-y
van Zoest, V., Varotsis, G., Menzel, U., Wigren, A., Kennedy, B., Martinell, M., Fall, T. (2022) Spatio-temporal predictions of COVID-19 test positivity in Uppsala County, Sweden: a comparative approach. Scientific Reports, 7, 15176.