Newly discovered Fu2 nanobody hold potential for future antiviral development
Published: 2022-02-17
The COVID-19 pandemic, which is going into its third year, has had major societal and economic consequences worldwide. Despite lockdowns and non-pharmaceutical interventions (NPIs), such as restrictions on gatherings, social distancing requirements, and face mask mandates, the pandemic is still ongoing. The vaccine roll-out is progressing, and several treatment options and drugs are now registered for use. However, the continuous emergence of new Variants of Concern (VoC) has delayed the re-opening of society. It is unfortunately likely to continue to do so, as research has confirmed that new variants are less effectively neutralised by antibodies elicited by the first-generation vaccines (cf Cele et al. 2021). The development of novel treatments and vaccines that protects against SARS-CoV-2 variants, particularly VoCs, is therefore urgently needed.
Previous research has suggested the use of camelid-derived single-domain antibody fragments, so-called ‘nanobodies’, as early-phase SARS-CoV-2 therapeutic agents (cf Hanke et al. 2020; Moliner-Morro et al. 2020; Koenig et al. 2021).
Nanobodies offer several advantages over other therapies; they are small, simple to clone and express, stable, have high specificity and affinity when binding antigens, and exhibit favorable biochemical characteristics, such as good thermal stability. See our previous data highlights here and here for more details. In addition, the production of nanobodies for therapies is highly scalable, and they can be mass produced relatively cost effectively. It is also possible to combine the nanobodies to form homodimers/heterodimers/multimers, to both increase their potency and target multiple epitopes at the same time. All of this makes them ideal for use as a treatment, as it unlikely to be at risk of being rendered less effective due to viral escape.
Research has, over the last few years, helped to elucidate how SARS-CoV-2 attaches to the host cell receptor (ACE2) using the viral spike (S) protein, and the mechanism enabling subsequent fusion. Studies have shown that SARS-CoV-2 can be easily neutralised by, for example, high-affinity binding antibodies and nanobodies that are binding directly or close to the receptor binding motif of the receptor binding domain (RBD), thereby neutralising the virus by preventing attachment. However, as new virus variants emerge that can, at least in part, evade antibody-mediated neutralization have emerged and warrants more research.
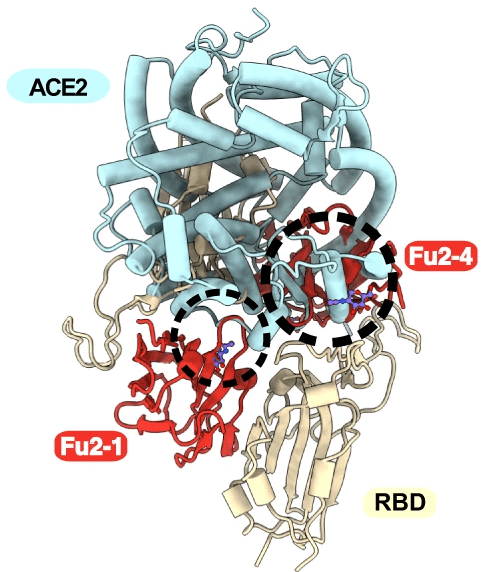
In a recently published article researchers from Karolinska Institutet (first authors: Leo Hanke, Hrishikesh Das; corresponding authors: B. Martin Hällberg (Karolinska Institutet/Centre for Stuctural Systems Biologi, Hamburg, Germany), Ben Murrell, and Gerald M. McInerney) together with colleagues describe how they generated a nanobody with two nonoverlapping interaction sites. The researchers showed that this new nanobody was able to interact with two RBDs from different spike trimers of SARS-CoV-2 at the same time. This interaction induced the formation of spike trimer–dimers, which caused SARS-CoV-2 to lose its ability to attach to host cell receptor ACE2.
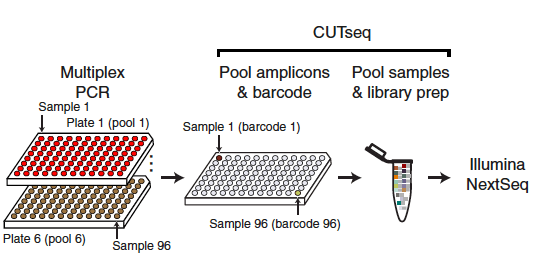
In order to identify new nanobodies with a unique mode of neutralization, the researchers first immunised an alpaca, with recombinant SARS-CoV-2 spike protein and RBD. After isolating RNA from peripheral blood mononuclear cells and creating phagemid libraries, phage display screens with immobilised spike protein were carried out. The phage display screens identified one nanobody of particular interest, named Fu2. In the next step, Hanke created nanobody multimers.
Previous research has indicated that multimerisation of nanobodies may be one way to improve their virus neutralization potency. Hanke and colleagues created three different dimers; an Fu2-Fc fusion expressed in mammalian cells, a chemically linked Fu2 homodimer, and a heterodimer of Fu2 with a previously described RBD-specific neutralising nanobody Ty1 Hanke et al 2020 (see our previous data highlight for details). The results also indicated that the Fu2 nanobody has the potential to both neutralise multiple VoCs (e.g. the Beta and Delta variants), and cross-neutralise SARS-CoV. Furthermore, the researchers also demonstrated the therapeutic potential of the nanobody against SARS-CoV-2 and the SARS-CoV-2 beta variant in a human ACE2 transgenic mouse model. Notably, for the Fu2-Ty1 heterodimer, no protection was found if it was administrated prophylactically, but therapeutic administration was found to significantly delay disease onset and disease severity. In summary, nanobodies could be a valuable alternative to monoclonal antibodies for use as antivirals against SARS-CoV-2 and other viruses.
The results by Hanke, Das, Sheward et al. show a naturally elicited bispecific monomeric nanobody active against both the original wild-type SARS-CoV-2 and VoCs. The RBD-specific nanobody Fu2 uses a novel strategy to neutralise SARS-CoV-2 by sterically blocking ACE2 interaction and inducing the formation of spike trimer–dimers which leads to aggregation of virions. Nanobodies with potentially new neutralization mechanisms that can inactivate viral antigens may be important for fighting both the current pandemic and future pandemics, making them potentially important for pandemic preparedness.
Data
- Cryo-EM density maps are deposited in the Electron Microscopy Data Bank under accession codes EMD-12561 (dimer of spike trimer + 6 Fu2) and EMD-12465 (localized reconstruction of 2 RBDs + 2 Fu2).
- The atomic coordinates have been deposited in the Protein Data Bank under IDs 7NS6 (dimers of spike trimers + 6 Fu2) and 7NLL (localized reconstruction of 2 RBDs + 2 Fu2) and will be available soon.
Article
DOI: 10.1038/s41467-021-27610-z
Hanke, L., Das, H., Sheward, D.J. et al. A bispecific monomeric nanobody induces spike trimer dimers and neutralizes SARS-CoV-2 in vivo. Nat Commun 13, 155 (2022).
Funding
The research was supported by the David och Astrid Hageléns stiftelse, the Clas Groschinskys Minnesfond and a Jonas Söderquist’s scholarship. In addition, the project has received funding from the European Union’s Horizon 2020 research and innovation program under grant agreement No. 101003653 (CoroNAb), Knut and Alice Wallenberg Foundation, Swedish Research Council, Karolinska Institutet Foundation Grants, National Molecular Medicine Program Grants, and the grants from the SciLifeLab National COVID-19 Research Program, financed by the Knut and Alice Wallenberg Foundation.