Multidrug‑resistant Enterobacterales in Swedish wastewater effluent
Published: 2026-03-30
Enterobacterales is an order of gram-negative bacteria that includes many pathogens. Carbapenem‑resistant Enterobacterales (CRE) carry genes that enable the production of carbapenemase, which breaks down carbapenem (a class of antibiotic agents). CRE, especially Klebsiella pneumoniae and Escherichia coli, represent major threats to public health; they cause severe opportunistic infections, and are often resistant to multiple classes of antibiotics.
In 2024, CRE suddenly appeared in outlet water from the Kristianstad wastewater treatment plant (WWTP). To investigate this, Axelsson et al. (2026) completed intense sampling of the outlet water for two weeks between September and October 2024. They subsequently obtained 55 CRE isolates. The Microflex Biotyper MALDI‑ToF MS (Bruker, Bremen, Germany) with the 1,829,023 Maldi Biotyper Compass Library was used to identify these isolates. In total, 29 isolates were identified as E. coli, and 26 as K. pneumoniae.
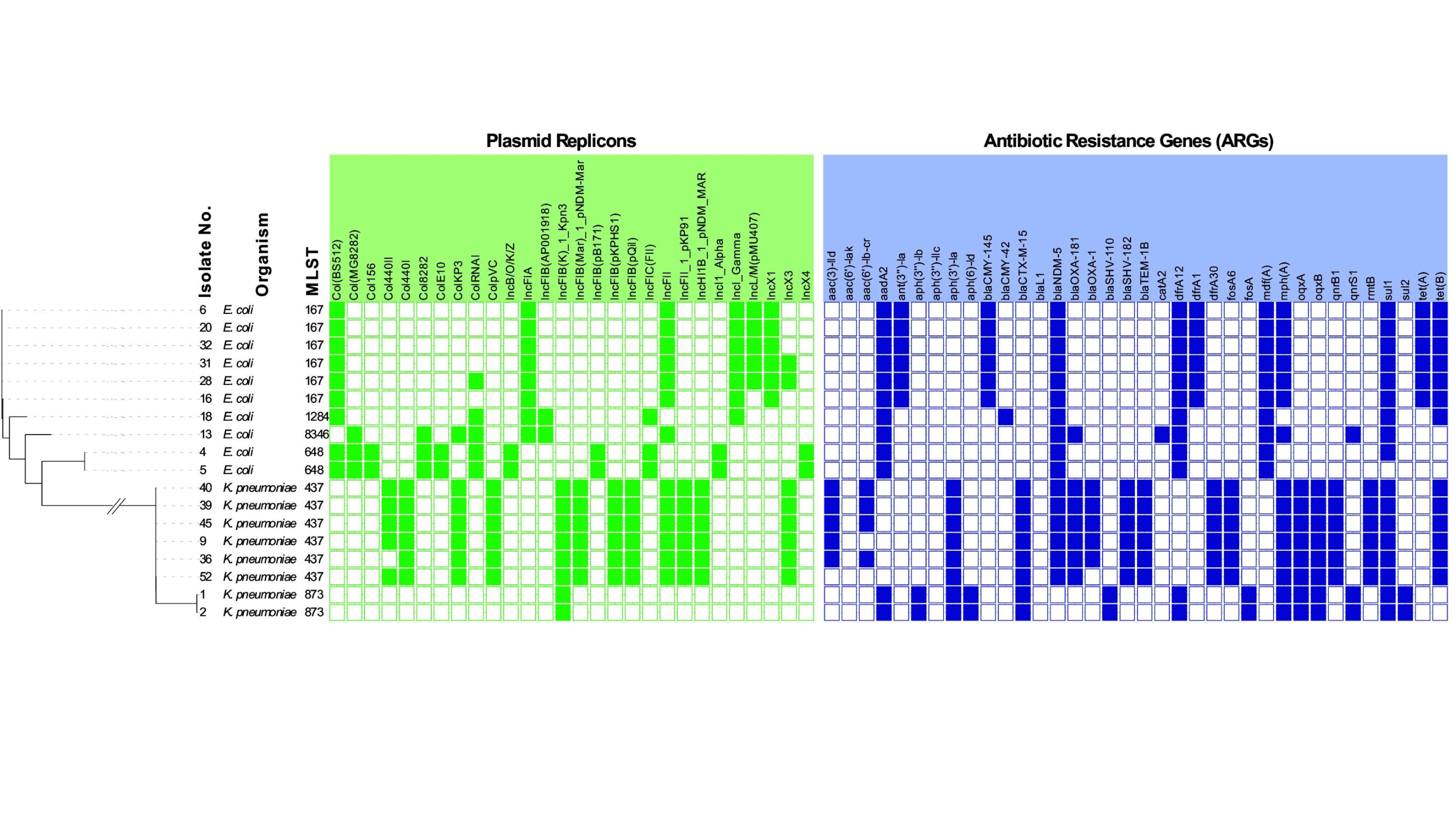
Real‑time PCR and melt‑curve analyses were done to assess the prevalence of genes related to carbapenemase production in the E. coli and K. pneumoniae isolates. The analyses revealed that 94% of the isolates carried blaNDM-5 genes and 56% carried blaOXA‑48 ‑ like genes. This confirmed widespread carbapenemase production in the effluent microbiota. Eighteen representative isolates (10 E. coli, and 8 K. pneumoniae) were selected for whole‑genome sequencing (WGS) to characterise sequence types, plasmids, virulence factors, and antibiotic resistance genes. Phylogenomic analysis showed that all of the sequenced isolates were multidrug‑resistant, with sequences exhibiting conserved plasmid profiles and extensive resistance genes.
The E. coli isolates belonged to globally important, high‑risk sequence types (STs) ST167, ST648, ST1284, and the emerging ST8346. K. pneumoniae isolates were from sequence types ST437 and ST873. ResFinder analysis showed blaNDM‑5 in all E. coli STs, and in K. pneumoniae ST437. Further, co‑carriage of blaNDM‑5 and blaOXA‑181, together with extended-spectrum-β-lactamases (ESBL) gene blaCTX‑M‑15, occurred in K. pneumoniae ST437 and E. coli ST8346. This indicates high levels of antibiotic resistance. Although K. pneumoniae ST437 carried virulence‑associated siderophore systems (e.g. yersiniabactin), it lacked the full hypervirulence, indicating classic, but still epidemiologically high‑risk, strains.
Two K. pneumoniae ST873 isolates from March 2024 were found to be carbapenem‑intermediate or carbapenem‑resistant, although they lacked known carbapenemase genes. They harboured only ESBL genes blaSHV‑110 and blaCTX‑M‑15, suggesting alternative or yet‑unidentified resistance mechanisms. The E. coli STs detected, including ST167, ST648, ST1284, and ST8346, are all associated with urinary tract, bloodstream, and wound infections, highlighting the clinical relevance of the wastewater isolates. Notably, ST8346 has recently been reported as an emerging clone with plasmid‑borne co‑carriage of blaNDM‑5 and blaOXA‑181, consistent with the genetic profiles observed here.
The Kristianstad wastewater treatment plant (WWTP) receives wastewater from approximately 210,000 people, the city hospital, and a large pig slaughterhouse. It had been subject to over a decade of surveillance (2014–2024) without detecting CRE before this event. The appearance of blaNDM‑5‑positive ST437 K. pneumoniae and ST167 E. coli in treated effluent therefore suggests a recent introduction and spread of high‑risk clones in the community that are not yet fully captured by clinical surveillance. Because WWTP inlets reflect carriage in the wider population and treated effluent can disseminate resistant bacteria into receiving waters, the findings imply an increased risk of environmental-, and potentially human-transmission of carbapenem‑resistant pathogens.
By linking wastewater‑based surveillance with genomic characterisation, Axelsson et al. (2026) demonstrates how high‑risk CRE lineages can silently establish in the environment, even in countries with relatively low clinical incidence. It underscores the need for integrated One Health monitoring of resistance genes across clinical, community, and environmental compartments, and highlights WWTP effluents as critical control points for limiting the spread of multidrug‑resistant Enterobacterales.
Data
Sequencing data is available at GenBank under reference numbers JBRMLF000000000 to JBRMLW000000000.
Article
DOI: 10.1017/s0950268826101071
Axelsson, C., de Neczpal, A. J., Sjöling, Å., & Rehnstam-Holm, A. S. (2026). Emergence of blaNDM-5 Enterobacterales in Swedish wastewater effluent. Epidemiology & Infection, 154, e23.